2024
|
 | Ouellet, Mathieu; Bassett, Dani S; Bassett, Lee C; Murphy, Kieran A; Patankar, Shubhankar P Mechanical prions: Self-assembling microstructures Journal Article Forthcoming Forthcoming. Abstract | Links | BibTeX | Tags: complex systems @article{Ouellet2024,
title = {Mechanical prions: Self-assembling microstructures},
author = {Mathieu Ouellet and Dani S. Bassett and Lee C. Bassett and Kieran A. Murphy and Shubhankar P. Patankar},
url = {https://arxiv.org/abs/2402.10939},
year = {2024},
date = {2024-04-16},
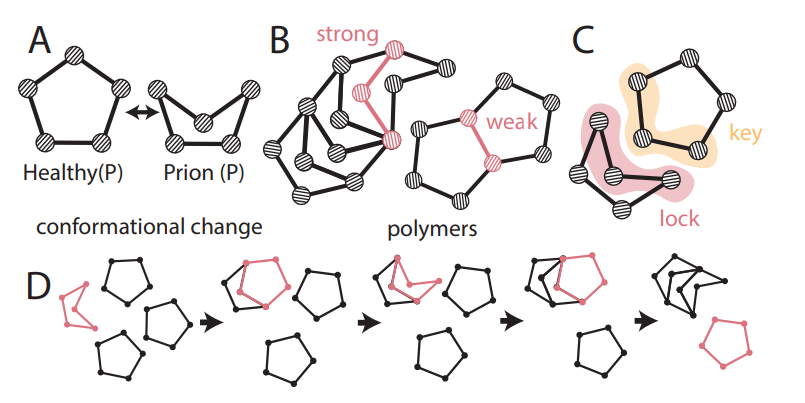
abstract = {Prions are misfolded proteins that transmit their structural arrangement to neighboring proteins. In biological systems, prion dynamics can produce a variety of complex functional outcomes. Yet, an understanding of prionic causes has been hampered by the fact that few computational models exist that allow for experimental design, hypothesis testing, and control. Here, we identify essential prionic properties and present a biologically inspired model of prions using simple mechanical structures capable of undergoing complex conformational change. We demonstrate the utility of our approach by designing a prototypical mechanical prion and validating its properties experimentally. Our work provides a design framework for harnessing and manipulating prionic properties in natural and artificial systems.},
keywords = {complex systems},
pubstate = {forthcoming},
tppubtype = {article}
}
Prions are misfolded proteins that transmit their structural arrangement to neighboring proteins. In biological systems, prion dynamics can produce a variety of complex functional outcomes. Yet, an understanding of prionic causes has been hampered by the fact that few computational models exist that allow for experimental design, hypothesis testing, and control. Here, we identify essential prionic properties and present a biologically inspired model of prions using simple mechanical structures capable of undergoing complex conformational change. We demonstrate the utility of our approach by designing a prototypical mechanical prion and validating its properties experimentally. Our work provides a design framework for harnessing and manipulating prionic properties in natural and artificial systems. |
 | Ouellet, Mathieu; Kim, Jason Z; Guillaume, Harmange; Shaffer, Sydney M; Bassett, Lee C; Bassett, Dani S Breaking reflection symmetry: evolving long dynamical cycles in Boolean systems Journal Article New Journal of Physics, 26 , pp. 023006, 2024. Abstract | Links | BibTeX | Tags: complex systems @article{Ouellet2024b,
title = {Breaking reflection symmetry: evolving long dynamical cycles in Boolean systems},
author = {Mathieu Ouellet and Jason Z Kim and Harmange Guillaume and Sydney M Shaffer and Lee C Bassett and Dani S Bassett},
url = {https://iopscience.iop.org/article/10.1088/1367-2630/ad1bdd/meta},
doi = {10.1088/1367-2630/ad1bdd},
year = {2024},
date = {2024-02-06},
journal = {New Journal of Physics},
volume = {26},
pages = {023006},
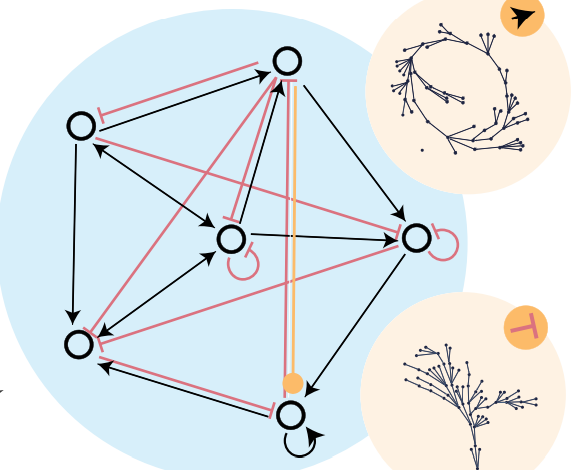
abstract = {In interacting dynamical systems, specific local interaction rules for system components give rise to diverse and complex global dynamics. Long dynamical cycles are a key feature of many natural interacting systems, especially in biology. Examples of dynamical cycles range from circadian rhythms regulating sleep to cell cycles regulating reproductive behavior. Despite the crucial role of cycles in nature, the properties of network structure that give rise to cycles still need to be better understood. Here, we use a Boolean interaction network model to study the relationships between network structure and cyclic dynamics. We identify particular structural motifs that support cycles, and other motifs that suppress them. More generally, we show that the presence of dynamical reflection symmetry in the interaction network enhances cyclic behavior. In simulating an artificial evolutionary process, we find that motifs that break reflection symmetry are discarded. We further show that dynamical reflection symmetries are over-represented in Boolean models of natural biological systems. Altogether, our results demonstrate a link between symmetry and functionality for interacting dynamical systems, and they provide evidence for symmetry's causal role in evolving dynamical functionality.},
keywords = {complex systems},
pubstate = {published},
tppubtype = {article}
}
In interacting dynamical systems, specific local interaction rules for system components give rise to diverse and complex global dynamics. Long dynamical cycles are a key feature of many natural interacting systems, especially in biology. Examples of dynamical cycles range from circadian rhythms regulating sleep to cell cycles regulating reproductive behavior. Despite the crucial role of cycles in nature, the properties of network structure that give rise to cycles still need to be better understood. Here, we use a Boolean interaction network model to study the relationships between network structure and cyclic dynamics. We identify particular structural motifs that support cycles, and other motifs that suppress them. More generally, we show that the presence of dynamical reflection symmetry in the interaction network enhances cyclic behavior. In simulating an artificial evolutionary process, we find that motifs that break reflection symmetry are discarded. We further show that dynamical reflection symmetries are over-represented in Boolean models of natural biological systems. Altogether, our results demonstrate a link between symmetry and functionality for interacting dynamical systems, and they provide evidence for symmetry's causal role in evolving dynamical functionality. |