Quantum Engineering Laboratory
Our group studies quantum dynamics in nanoscale materials and devices using optics and electronics. We seek to better understand complex quantum-mechanical systems, with a goal of developing new technologies for communication, computation, and sensing based on quantum physics.









News
- Congratulations, Sarah!
Sarah has won the Haller Prize, awarded to the best graduate student at the 32nd International Conference on Defects in Semiconductors. “The Haller Prize is named after Eugene E. Haller who was a major figure in the semiconductor community and an inspiring mentor for students.” (http://icds2023.org/prizes)
- Welcome, Jeiko!
Jeiko Pujols has joined our group as a first-year PhD student in the ESE department.
- Congratulations, Henry!
Congratulations to Henry on defending his PhD thesis!

- Congratulations, Alex!
Congratulations to Alex on defending his PhD thesis!

- Congratulations, Raj!
Congratulations to Raj on defending his PhD thesis!

Recent Publications
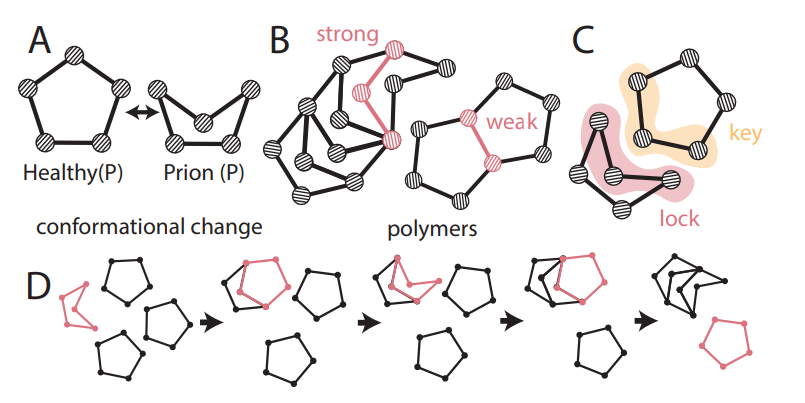 | Ouellet, Mathieu; Bassett, Dani S; Bassett, Lee C; Murphy, Kieran A; Patankar, Shubhankar P Mechanical prions: Self-assembling microstructures Journal Article Forthcoming Forthcoming. @article{Ouellet2024, title = {Mechanical prions: Self-assembling microstructures}, author = {Mathieu Ouellet and Dani S. Bassett and Lee C. Bassett and Kieran A. Murphy and Shubhankar P. Patankar}, url = {https://arxiv.org/abs/2402.10939}, year = {2024}, date = {2024-04-16}, abstract = {Prions are misfolded proteins that transmit their structural arrangement to neighboring proteins. In biological systems, prion dynamics can produce a variety of complex functional outcomes. Yet, an understanding of prionic causes has been hampered by the fact that few computational models exist that allow for experimental design, hypothesis testing, and control. Here, we identify essential prionic properties and present a biologically inspired model of prions using simple mechanical structures capable of undergoing complex conformational change. We demonstrate the utility of our approach by designing a prototypical mechanical prion and validating its properties experimentally. Our work provides a design framework for harnessing and manipulating prionic properties in natural and artificial systems.}, keywords = {}, pubstate = {forthcoming}, tppubtype = {article} } Prions are misfolded proteins that transmit their structural arrangement to neighboring proteins. In biological systems, prion dynamics can produce a variety of complex functional outcomes. Yet, an understanding of prionic causes has been hampered by the fact that few computational models exist that allow for experimental design, hypothesis testing, and control. Here, we identify essential prionic properties and present a biologically inspired model of prions using simple mechanical structures capable of undergoing complex conformational change. We demonstrate the utility of our approach by designing a prototypical mechanical prion and validating its properties experimentally. Our work provides a design framework for harnessing and manipulating prionic properties in natural and artificial systems. |
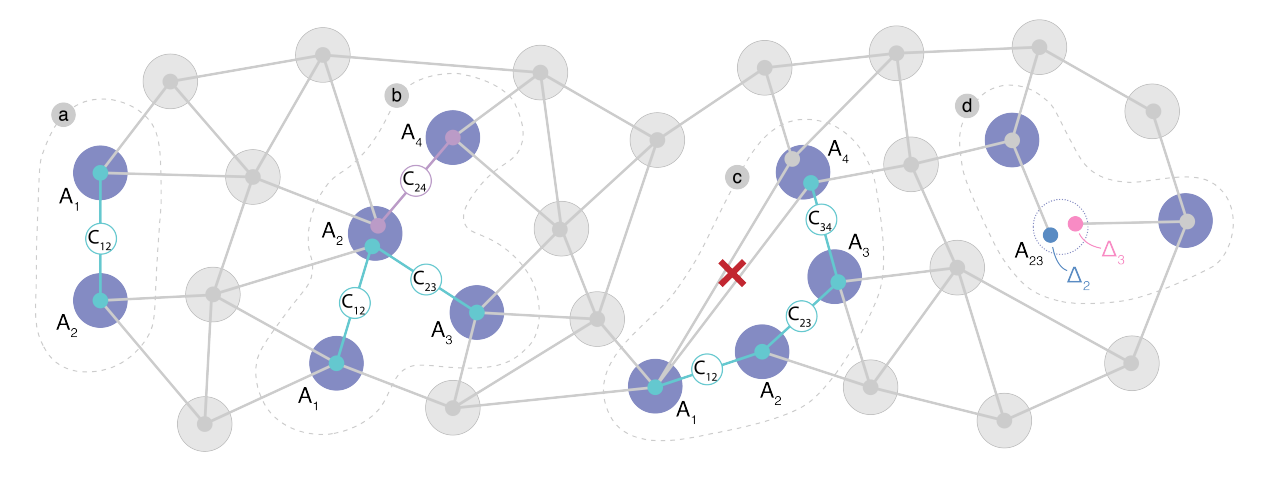 | van de Stolpe, G L; Kwiatkowski, D P; Bradley, C E; Randall, J; Breitweiser, S A; Bassett, L C; Markham, M; Twitchen, D J; Taminiau, T H Mapping a 50-spin-qubit network through correlated sensing Journal Article Nature Communications , 15 (2006), 2024. @article{vandeStolpe2023, title = {Mapping a 50-spin-qubit network through correlated sensing}, author = {G.L. van de Stolpe and D. P. Kwiatkowski and C.E. Bradley and J. Randall and S. A. Breitweiser and L. C. Bassett and M. Markham and D.J. Twitchen and T.H. Taminiau}, url = {https://arxiv.org/abs/2307.06939 https://www.nature.com/articles/s41467-024-46075-4}, doi = {10.1038/s41467-024-46075-4}, year = {2024}, date = {2024-03-05}, journal = {Nature Communications }, volume = {15}, number = {2006}, abstract = {Spins associated to optically accessible solid-state defects have emerged as a versatile platform for exploring quantum simulation, quantum sensing and quantum communication. Pioneering experiments have shown the sensing, imaging, and control of multiple nuclear spins surrounding a single electron-spin defect. However, the accessible size and complexity of these spin networks has been constrained by the spectral resolution of current methods. Here, we map a network of 50 coupled spins through high-resolution correlated sensing schemes, using a single nitrogen-vacancy center in diamond. We develop concatenated double-resonance sequences that identify spin-chains through the network. These chains reveal the characteristic spin frequencies and their interconnections with high spectral resolution, and can be fused together to map out the network. Our results provide new opportunities for quantum simulations by increasing the number of available spin qubits. Additionally, our methods might find applications in nano-scale imaging of complex spin systems external to the host crystal.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Spins associated to optically accessible solid-state defects have emerged as a versatile platform for exploring quantum simulation, quantum sensing and quantum communication. Pioneering experiments have shown the sensing, imaging, and control of multiple nuclear spins surrounding a single electron-spin defect. However, the accessible size and complexity of these spin networks has been constrained by the spectral resolution of current methods. Here, we map a network of 50 coupled spins through high-resolution correlated sensing schemes, using a single nitrogen-vacancy center in diamond. We develop concatenated double-resonance sequences that identify spin-chains through the network. These chains reveal the characteristic spin frequencies and their interconnections with high spectral resolution, and can be fused together to map out the network. Our results provide new opportunities for quantum simulations by increasing the number of available spin qubits. Additionally, our methods might find applications in nano-scale imaging of complex spin systems external to the host crystal. |
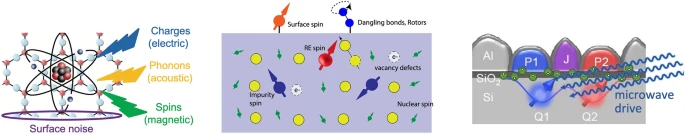 | Gali, Adam; Schleife, André; Heinrich, Andreas J; Laucht, Arne; Schuler, Bruno; Chakraborty, Chitraleema; Anderson, Christopher P; Déprez, Corentin; McCallum, Jeffrey; Bassett, Lee C; Friesen, Mark; Flatté, Michael E; Maurer, Peter; Coppersmith, Susan N; Zhong, Tian; Begum-Hudde, Vijaya; Ping, Yuan Challenges in advancing our understanding of atomic-like quantum systems: Theory and experiment Journal Article MRS Bulletin, 49 , pp. 256-276, 2024. @article{Gali2024, title = {Challenges in advancing our understanding of atomic-like quantum systems: Theory and experiment}, author = {Adam Gali and André Schleife and Andreas J. Heinrich and Arne Laucht and Bruno Schuler and Chitraleema Chakraborty and Christopher P. Anderson and Corentin Déprez and Jeffrey McCallum and Lee C. Bassett and Mark Friesen and Michael E. Flatté and Peter Maurer and Susan N. Coppersmith and Tian Zhong and Vijaya Begum-Hudde and Yuan Ping }, url = {https://link.springer.com/article/10.1557/s43577-023-00659-5}, doi = {10.1557/s43577-023-00659-5}, year = {2024}, date = {2024-02-14}, journal = {MRS Bulletin}, volume = {49}, pages = {256-276}, abstract = {Quantum information processing and quantum sensing is a central topic for researchers who are part of the Materials Research Society and the Quantum Staging Group is providing leadership and guidance in this context. We convened a workshop before the 2022 MRS Spring Meeting and covered four topics to explore challenges that need to be addressed to further promote and accelerate the development of materials with applications in quantum technologies. This article captures the discussions at this workshop and refers to the pertinent literature.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Quantum information processing and quantum sensing is a central topic for researchers who are part of the Materials Research Society and the Quantum Staging Group is providing leadership and guidance in this context. We convened a workshop before the 2022 MRS Spring Meeting and covered four topics to explore challenges that need to be addressed to further promote and accelerate the development of materials with applications in quantum technologies. This article captures the discussions at this workshop and refers to the pertinent literature. |
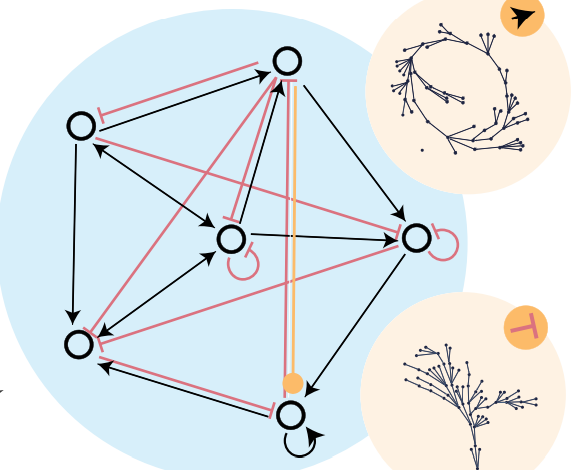 | Ouellet, Mathieu; Kim, Jason Z; Guillaume, Harmange; Shaffer, Sydney M; Bassett, Lee C; Bassett, Dani S Breaking reflection symmetry: evolving long dynamical cycles in Boolean systems Journal Article New Journal of Physics, 26 , pp. 023006, 2024. @article{Ouellet2024b, title = {Breaking reflection symmetry: evolving long dynamical cycles in Boolean systems}, author = {Mathieu Ouellet and Jason Z Kim and Harmange Guillaume and Sydney M Shaffer and Lee C Bassett and Dani S Bassett}, url = {https://iopscience.iop.org/article/10.1088/1367-2630/ad1bdd/meta}, doi = {10.1088/1367-2630/ad1bdd}, year = {2024}, date = {2024-02-06}, journal = {New Journal of Physics}, volume = {26}, pages = {023006}, abstract = {In interacting dynamical systems, specific local interaction rules for system components give rise to diverse and complex global dynamics. Long dynamical cycles are a key feature of many natural interacting systems, especially in biology. Examples of dynamical cycles range from circadian rhythms regulating sleep to cell cycles regulating reproductive behavior. Despite the crucial role of cycles in nature, the properties of network structure that give rise to cycles still need to be better understood. Here, we use a Boolean interaction network model to study the relationships between network structure and cyclic dynamics. We identify particular structural motifs that support cycles, and other motifs that suppress them. More generally, we show that the presence of dynamical reflection symmetry in the interaction network enhances cyclic behavior. In simulating an artificial evolutionary process, we find that motifs that break reflection symmetry are discarded. We further show that dynamical reflection symmetries are over-represented in Boolean models of natural biological systems. Altogether, our results demonstrate a link between symmetry and functionality for interacting dynamical systems, and they provide evidence for symmetry's causal role in evolving dynamical functionality.}, keywords = {}, pubstate = {published}, tppubtype = {article} } In interacting dynamical systems, specific local interaction rules for system components give rise to diverse and complex global dynamics. Long dynamical cycles are a key feature of many natural interacting systems, especially in biology. Examples of dynamical cycles range from circadian rhythms regulating sleep to cell cycles regulating reproductive behavior. Despite the crucial role of cycles in nature, the properties of network structure that give rise to cycles still need to be better understood. Here, we use a Boolean interaction network model to study the relationships between network structure and cyclic dynamics. We identify particular structural motifs that support cycles, and other motifs that suppress them. More generally, we show that the presence of dynamical reflection symmetry in the interaction network enhances cyclic behavior. In simulating an artificial evolutionary process, we find that motifs that break reflection symmetry are discarded. We further show that dynamical reflection symmetries are over-represented in Boolean models of natural biological systems. Altogether, our results demonstrate a link between symmetry and functionality for interacting dynamical systems, and they provide evidence for symmetry's causal role in evolving dynamical functionality. |
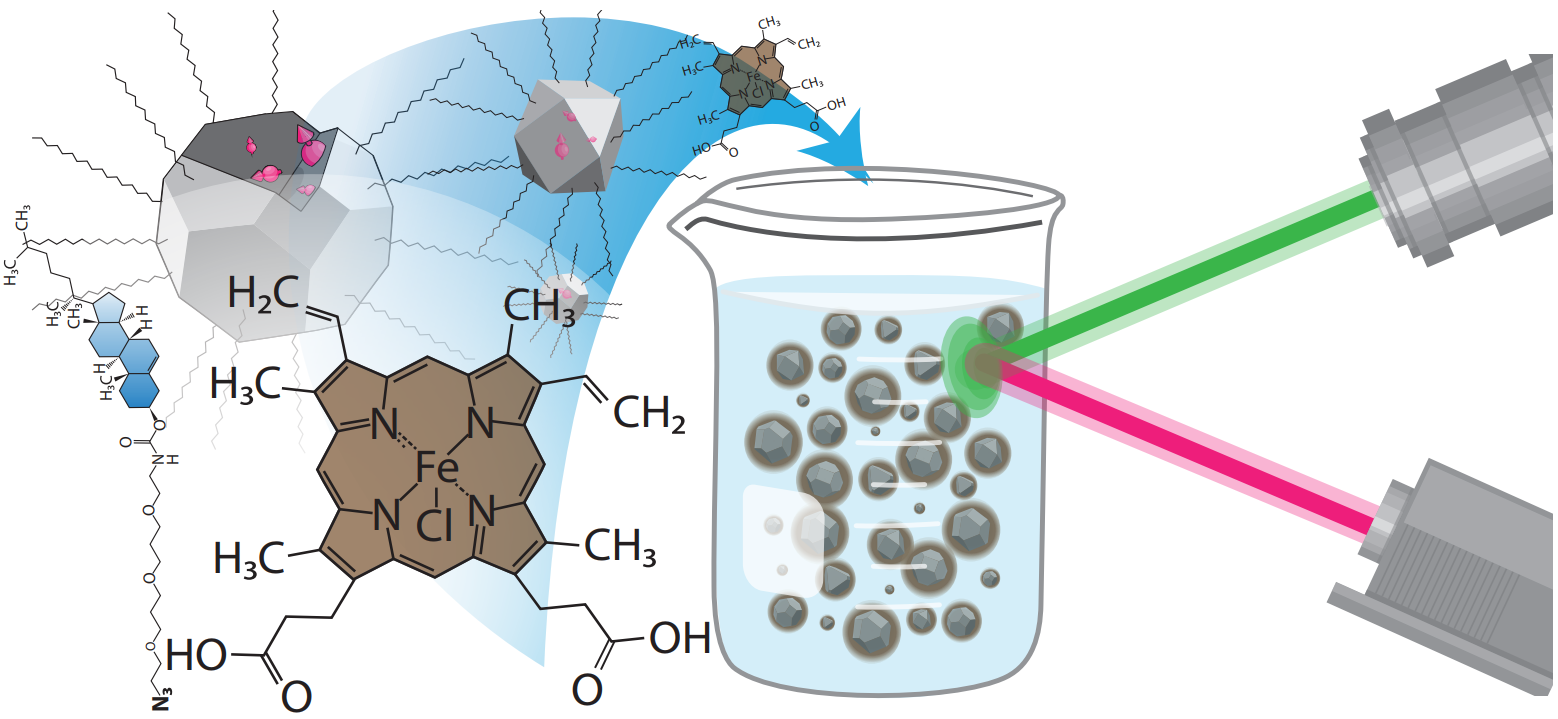 | Shulevitz, Henry J; Amirshaghaghi, Ahmad; Ouellet, Mathieu; Brustoloni, Caroline; Yang, Shengsong; Ng, Jonah J; Huang, Tzu-Yung; Jishkariani, Davit; Murray, Christopher B; Tsourkas, Andrew; Kagan, Cherie R; Bassett, Lee C Nanodiamond emulsions for enhanced quantum sensing and click-chemistry conjugation Journal Article Forthcoming Forthcoming. @article{Shulevitz2023, title = {Nanodiamond emulsions for enhanced quantum sensing and click-chemistry conjugation}, author = {Henry J. Shulevitz and Ahmad Amirshaghaghi and Mathieu Ouellet and Caroline Brustoloni and Shengsong Yang and Jonah J. Ng and Tzu-Yung Huang and Davit Jishkariani and Christopher B. Murray and Andrew Tsourkas and Cherie R. Kagan and Lee C. Bassett}, url = {https://arxiv.org/abs/2311.16530}, doi = {10.48550/arXiv.2311.16530}, year = {2023}, date = {2023-12-04}, abstract = {Nanodiamonds containing nitrogen-vacancy (NV) centers can serve as colloidal quantum sensors of local fields in biological and chemical environments. However, nanodiamond surfaces are challenging to modify without degrading their colloidal stability or the NV center's optical and spin properties. Here, we report a simple and general method to coat nanodiamonds with a thin emulsion layer that preserves their quantum features, enhances their colloidal stability, and provides functional groups for subsequent crosslinking and click-chemistry conjugation reactions. To demonstrate this technique, we decorate the nanodiamonds with combinations of carboxyl- and azide-terminated amphiphiles that enable conjugation using two different strategies. We study the effect of the emulsion layer on the NV center's spin lifetime, and we quantify the nanodiamonds' chemical sensitivity to paramagnetic ions using T1 relaxometry. This general approach to nanodiamond surface functionalization will enable advances in quantum nanomedicine and biological sensing.}, keywords = {}, pubstate = {forthcoming}, tppubtype = {article} } Nanodiamonds containing nitrogen-vacancy (NV) centers can serve as colloidal quantum sensors of local fields in biological and chemical environments. However, nanodiamond surfaces are challenging to modify without degrading their colloidal stability or the NV center's optical and spin properties. Here, we report a simple and general method to coat nanodiamonds with a thin emulsion layer that preserves their quantum features, enhances their colloidal stability, and provides functional groups for subsequent crosslinking and click-chemistry conjugation reactions. To demonstrate this technique, we decorate the nanodiamonds with combinations of carboxyl- and azide-terminated amphiphiles that enable conjugation using two different strategies. We study the effect of the emulsion layer on the NV center's spin lifetime, and we quantify the nanodiamonds' chemical sensitivity to paramagnetic ions using T1 relaxometry. This general approach to nanodiamond surface functionalization will enable advances in quantum nanomedicine and biological sensing. |